MSGIP – Mean Shift Genomic Island Preditor
Drito, DM; Maracaja-Coutinho, V; Farias, ST; Batista, LV; Rego, TG. (2016). A novel method to predict genomic islands based on mean shift clustering algorithm. PLOS One. DOI: 10.1371/journal.pone.0146352
MSGIP is an automated tool for the prediction of genomic islands based on mean shift algorithm. MSGIP was developed in Java and is compatible with any operation system with Java Runtime Environment installed. The usage requires only a FASTA “.fna” file, containing the complete genomic sequence of the investigated bacteria
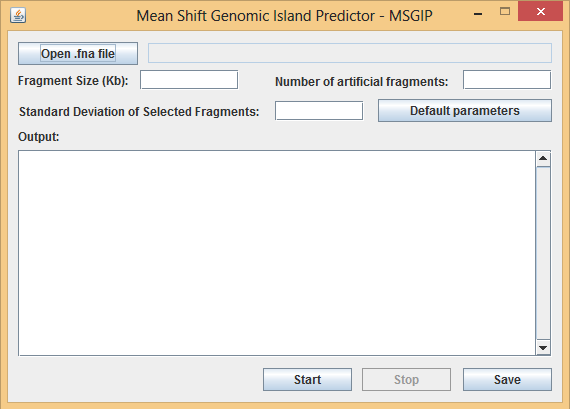 Figure 1 – Initial screen for the user-friendly version of MSGIP tool. The user define the parameters described on the manuscript from Brito et al. and upload the genomic sequence in FASTA format (.fna) in order to predict all potential genomic islands based on Mean Shift algorithm methodology.
Figure 1 – Initial screen for the user-friendly version of MSGIP tool. The user define the parameters described on the manuscript from Brito et al. and upload the genomic sequence in FASTA format (.fna) in order to predict all potential genomic islands based on Mean Shift algorithm methodology.
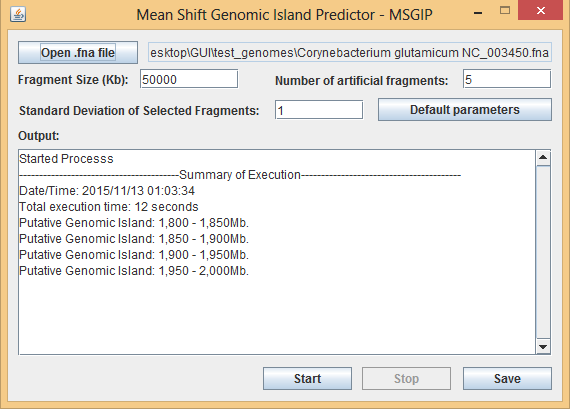 Figure 2 – Results screen for the user-friendly version of MSGIP tool. The predicted genomic islands are listed together with its genomic position on the sequence. The result can be saved and exported to a .txt file by clicking on the Save button.
Figure 2 – Results screen for the user-friendly version of MSGIP tool. The predicted genomic islands are listed together with its genomic position on the sequence. The result can be saved and exported to a .txt file by clicking on the Save button.
Downloading and Executing
MSGIP source code and user-friendly version are freely available and be can downloaded below:
- User-friendly version (1.0)
- Stand-alone version (1.0)
After downloading, please see the “read me” file provided.
You can also find MSGIP in our GitHub account. All information to install and run it can also be accessed there. Any questions please feel free to contact as on the e-mail provided on the end of this page.
Pre-Requisites
Contact
Please, feel free to contact us in case of comments, criticisms and suggestions at: britmb AT gmail DOT com

